Research overview
‘To suppose that the evolution of the wonderfully adapted biological mechanisms has depended only on a selection out of a haphazard set of variations, each produced by blind chance, is like suggesting that if we went on throwing bricks together into heaps, we should eventually be able to choose ourselves the most desirable house.’
- C. H. Waddington
Chromatin: a fundamental architectural component of eukaryotic genomes
The eukaryotic genome is compacted into the cell nucleus forming a nucleoprotein complex named chromatin. The fundamental unit of chromatin is the nucleosome, composed of 145-147 base pairs of genomic DNA wrapped around an octamer of core histones (H2A, H2B, H3 and H4). Chromatin is divided into two main functionally distinct domains with different DNA, RNA and protein composition. Euchromatin constitutes an open, accessible form of the chromatin and contains most actively expressed genes. On the other hand, heterochromatin harbours less accessible and often transcriptionally repressed or silent genes.
Distinct groups of writers, readers and erasers act together to impose particular histone post-translational modifications patterns (together known as the histone code) that define these highly conserved chromatin states and play a role in crucial biological processes such as DNA replication or transcription.
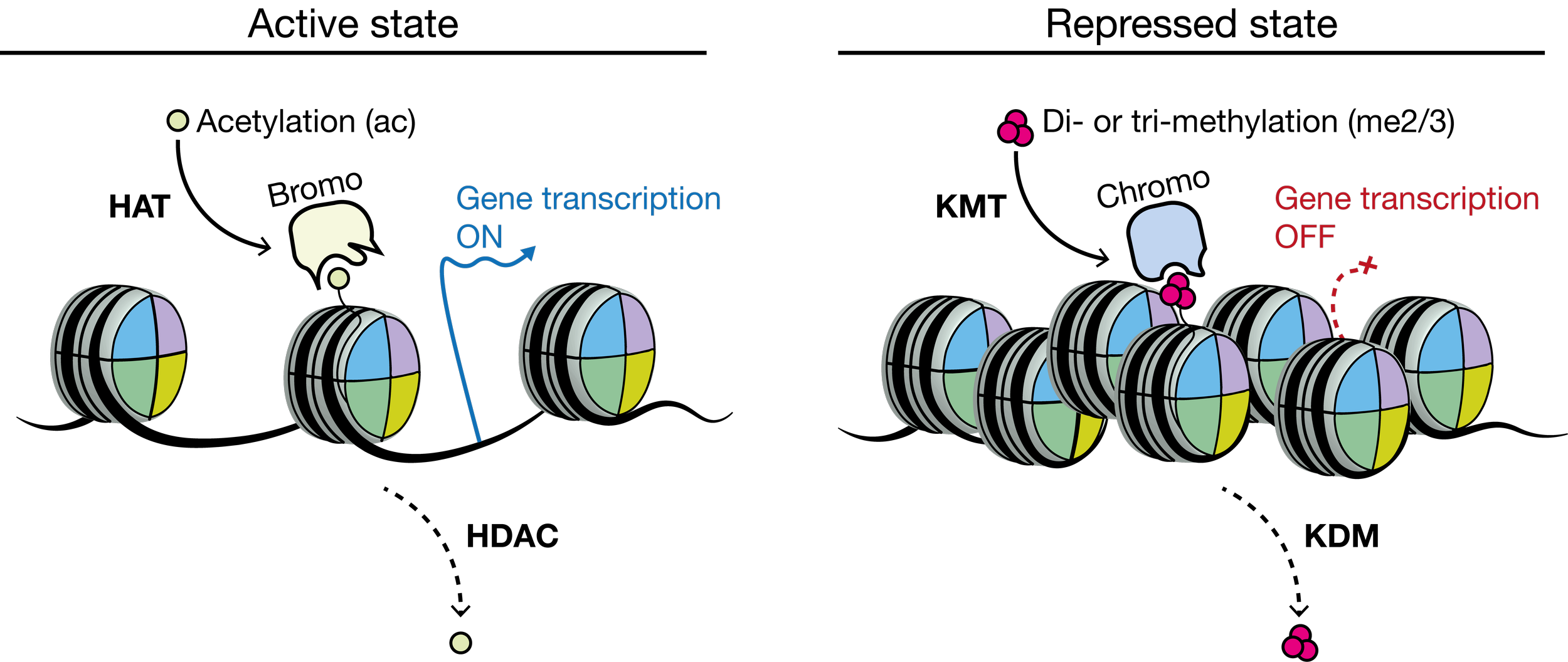
For example, acetylation of lysine residues by histone acetyltransferases (HATs) provides a docking site for bromodomain-containing proteins that bind to further stimulate nucleosome accessibility and transcriptional activity. Histone acetylation can be reversed by opposing histone deacetylases (HDACs), which often cause transcriptional repression. In contrast, lysine methyltransferases (KMTs) methylate the histone H3 amino-terminal tail (e.g. H3K9me). Tri-methylated H3K9 (H3K9me3) serves as a docking site for chromodomain-containing proteins, which then impede nucleosome accessibility and induce gene repression. Histone methylation can be reversed by opposing histone demethylases (KDMs).

Chromatin writers, readers and erasers. Distinct groups of proteins act together to impose particular histone modifications patterns that define transcriptionally active (left) or repressed (right) chromatin states. Adapted from Allis & Jenuwein (2016).
Adaptive epigenetic changes as a means of survival
But…what is epigenetics? Conrad Waddington coined the term epigenetics in the 1940s to refer to “the branch of biology that studies the causal interactions between genes and their products which bring the phenotype into being”. Robin Holliday redefined the term, formulating the concept of epimutation for the first time to describe “heritable changes in gene function that do not involve a change in DNA sequence”.
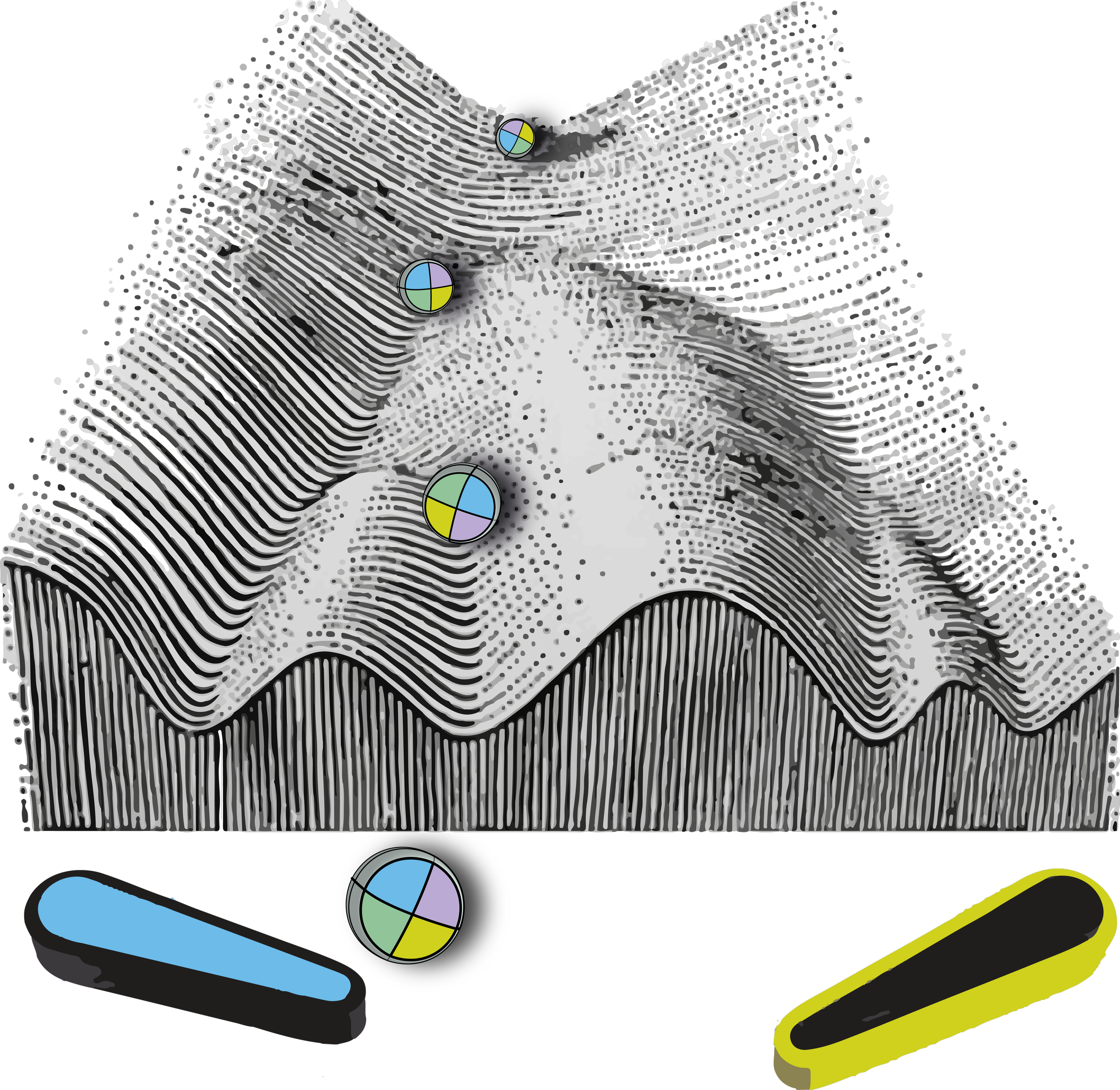
Changes in DNA sequence (mutations) that drive evolution are slow and nearly irreversible, and thus not ideal for a population to survive in a rapidly changing environment. Epimutations have been proposed to enable ‘soft inheritance’, resulting in the formation of different phenotypes that enable adaptation to fluctuating environments. This bet-hedging strategy would allow cells to adapt transiently to insults while remaining genetically wild-type, and thus the reversal of epimutations when selective pressures are relaxed.
During my PhD in Prof. Robin Allshire’s laboratory at the Wellcome Centre for Cell Biology, University of Edinburgh I showed that heterochromatin-mediated epimutations can drive reversible drug resistance in fission yeast. This proof-of-concept study provided foundational evidence for chromatin-based epialleles with adaptive impact.
Currently, I investigate whether similar mechanisms can play a role in the emergence and evolution of therapy resistance in metastatic cancer – a major biomedical challenge wherein tumor cells from a primary site gradually colonize distant organs. Understanding the contribution of epigenetic processes to cancer progression under therapy could shed light on conserved principles of eukaryotic adaptation and reveal potential therapeutic opportunities to improve patient outcome.